1. Introduction
Emory University’s High Performance Computing (HPC) cluster provides researchers with powerful computational resources for running large-scale analyses. This tutorial demonstrates how to run simulation studies in R using the HPC’s job scheduling system (SLURM). For more details, visit the RSCH HPC Documentation.
🌱Project Overview
The example project investigates logistic regression vs. linear regression performance on binary outcome data across different sample sizes. Key parameters:
- Sample sizes: 500, 1000, 2000
- Models: Logistic regression, Linear regression
- Replications: 1,000 per scenario
- Total jobs: 6,000 (2 models × 3 sample sizes × 1,000 replications)
2. Local R Project
First, create an R project on your local computer. Organize your files so that the HPC folder contains jobs that will be submitted to the cluster and receive results from it. The complete code is available on GitHub: xxou
Emory-HPC.
‼️NOTE: We use the here() package to ensure consistent working directory paths across local and HPC environments.
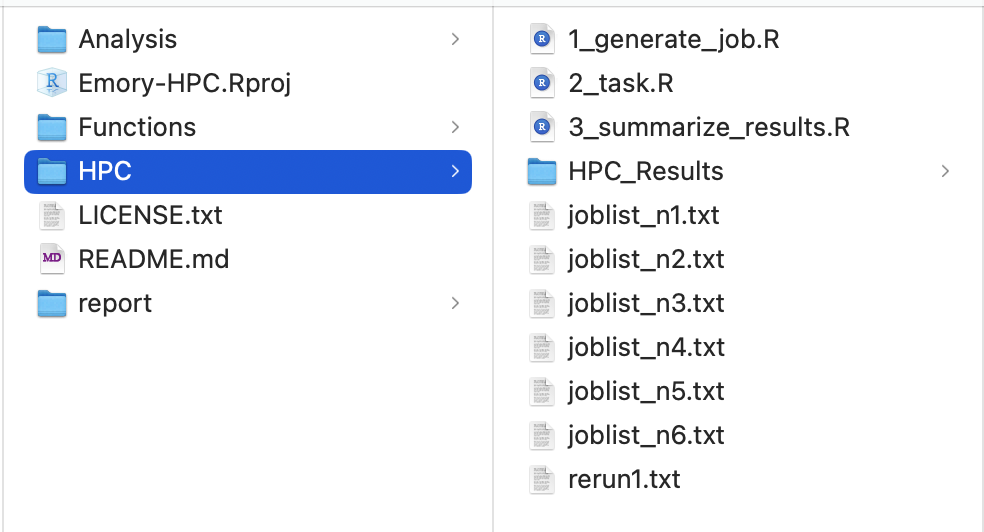
2.1 Generate Simulation Jobs
The file 1_generate_job.R generates simulation job lists. It creates several text files containing commands that will run 2_task.R on the HPC.
# *****************************************************
# 1. Generate simulation jobs
# *****************************************************
# Number of simulation replicates per scenario
sims = 1000
# Generate parameter grid: all combinations of seed, sample size, and model
param_grid <- expand.grid(seed = 1:sims,
n = c(500, 1000, 2000),
model = c("logistic","lm")
)
# Create job list: command to run each simulation task
# Format: "Rscript 2_task.R <seed> <n> <model>"
param_grid$joblist<-paste("Rscript 2_task.R",
param_grid$seed,
param_grid$n,
param_grid$model)
# Emory HPC limits: maximum 1000 jobs per job array chunk
# Split total jobs into chunks of at most 1000 jobs each
n_list <- nrow(param_grid) / sims
# Generate job list files (one per chunk)
for (i in 1:n_list) {
# Calculate row indices for the i-th chunk
start_row <- ((i - 1) * sims) + 1
end_row <- i * sims
# Write job list to a text file for HPC submission
write.table(param_grid[start_row:end_row, 'joblist'],
file =here::here("HPC/", paste0("joblist_n",i,".txt")),
row.names = FALSE, col.names = FALSE, quote = F)
}
Important: Emory HPC limits job arrays to 1,000 jobs per chunk. After running this code locally, you will have
joblist_n1.txtthroughjoblist_n6.txt.
2.2 Task File
The file 2_task.R runs the actual simulation. It will be executed on the cluster (not your local computer).
# ******************************************
# 2. Main task to run simulation
# ******************************************
freshr::freshr()
# Parse command-line arguments: seed, sample size, and model type
args = commandArgs(trailingOnly=T)
seed = as.numeric(args[1])
n = as.numeric(args[2])
model = as.character(args[3])
# Load required packages
pacman::p_load(tibble)
# Load data generation function
source(here::here("Functions/1_data_generation.R"))
# Generate synthetic data
set.seed(seed)
data <- DGP(n = n)
# Fit model based on specified type
if(model == "logistic"){
fit <- glm(y ~ x, data, family = binomial())
}
if(model == "lm"){
fit <- glm(y ~ x, data, family = gaussian())
}
# Create results table with estimated coefficients and scenario info
tbl <- tibble(
# Estimated coefficients
beta0 = coef(fit)[["(Intercept)"]],
beta1 = coef(fit)[["x"]],
# Scenario parameters
seed = seed,
n = n,
model = model
)
# Save results to file
save(tbl, file = here::here("HPC/HPC_Results", paste0("seed", seed, "_n", n, "_", model,".Rdata")))
Each task generates output stored in HPC/HPC_Results/.
2.3 Summarize Results
The file 3_summarize_results.R collects all results into a single file and identifies any failed jobs for rerunning.
# ******************************************
# 3. Summarize simulation results
# ******************************************
sims = 1000
# Reconstruct parameter grid to match job specifications
param_grid <- expand.grid(seed = 1:sims,
n = c(500, 1000, 2000),
model = c("logistic","lm")
)
# Construct expected output filenames
param_grid$file_name <-paste0("seed", param_grid$seed,
"_n", param_grid$n,
"_", param_grid$model,
".Rdata")
# For failed jobs: create rerun commands
param_grid$rerun_name<-paste("Rscript 2_task.R",
param_grid$seed,
param_grid$n,
param_grid$model)
# Load all results, handling potential job failures
AllResults<- list()
false_index = c()
for (i in 1:dim(param_grid)[1]) {
file_name = param_grid$file_name[i]
# Attempt to load the data file
load_status <- tryCatch({
load(here::here("HPC/HPC_Results", file_name))
TRUE
}, error = function(e) {
FALSE
})
if(load_status==F){
false_index = c(false_index,i)
}
if (load_status) {
AllResults[[i]] = tbl
}
}
# Save combined results
save(AllResults, file = here::here("HPC/HPC_Results", "AllResults.Rdata"))
# Write list of failed jobs for rerunning
write.table(unique(param_grid[false_index, 'rerun_name']),
file = here::here("HPC", "rerun1.txt"),
row.names = FALSE, col.names = FALSE, quote = F)
After preparing everything locally, let’s move to the HPC cluster.
3. Emory HPC Cluster
3.1 SSH Connection
Open your terminal, log in, and navigate to your project folder.
ssh your_emory_id@clogin01.sph.emory.edu
🌟First-time Users: Download dSQ (a job submission tool) to your HPC working directory.
git clone https://github.com/ycrc/dSQ
It shows in the terminal:
3.2 Transfer Files
Transfer your local project folder to HPC. Keep the folder structure unchanged—transfer at minimum: .Rproj, HPC folder, and related folder.
Option 1: Using scp
scp -r /local/path/to/Emory-HPC your_emory_id@clogin01.sph.emory.edu:~/path/to/destination/
Option 2: Using FileZilla
Download FileZilla from filezilla-project.org. This allows drag-and-drop file transfer. Enter the following in the quick connect bar:
- Host:
clogin01.sph.emory.edu - Username: Your Emory ID
- Password: Your Emory password
- Port: 22

3.3 Submit Your Job
Now you should have the same project folder structure on the cluster. Navigate to your HPC folder where the joblist_n*.txt files are located.
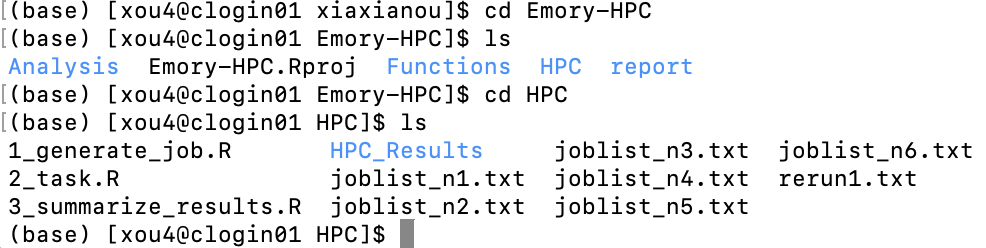
(1) Set up environment
Set the dSQ path (adjust to your actual path where you just downloaded dSQ):
dsq_path="/path/dSQ"
module load python/3.8
module load R/4.2.2
(2) Generate .sh files
for i in $(seq 1 6); do
python "$dsq_path/dSQ.py" --job-file joblist_n$i.txt --batch-file joblist_n$i.sh --job-name reg$i --mail-type ALL
done
for i in $(seq 1 6); do
sed -i 's@'"$dsq_path"'/dSQBatch.py@python '"$dsq_path"'/dSQBatch.py@' "joblist_n$i.sh"
sed -i '/#SBATCH --mail-type "ALL"/a #SBATCH --mail-user=your_email@emory.edu\nmodule load python/3.8\nmodule load R/4.2.2' joblist_n$i.sh
sed -i '1a #SBATCH --partition=short-cpu' joblist_n$i.sh
done
The first command generates .sh files from joblist_n*.txt. The second command customizes settings:
--job-name: Job identifier (currentlyreg)--mail-user: Your Emory email for status notifications--partition: Choose from the following: e.g. short-cpu (8 hours)
| Partition | Time Limit |
|---|---|
| short-cpu | 8:00:00 |
| day-long-cpu | 24:00:00 |
| week-long-cpu | 7-00:00:00 |
| month-long-cpu | 31-00:00:00 |
| interactive-cpu | 2:00:00 |
| largemem | 7-01:00:00 |
This generates 6 .sh files.
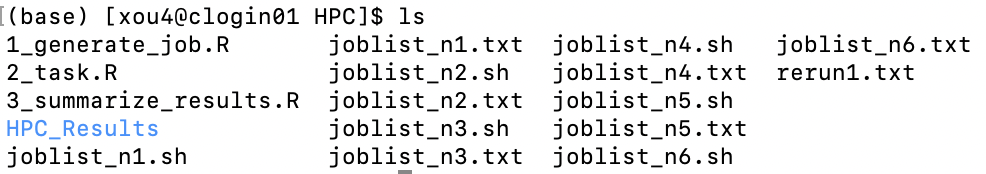
(3) Submit .sh files
Submit one job:
sbatch joblist_n1.sh
Submit all 6 jobs together:
for i in $(seq 1 6); do
sbatch joblist_n$i.sh
done
(4) Check job status
squeue --job <your_job_number> # Check specific job
squeue -u <your_username> # Check all your jobs
Cancel jobs:
scancel <your_job_number> # Cancel specific job
scancel <job_id1>,<job_id2>,<job_id3> # Cancel multiple jobs
scancel -u <your_username> # Cancel all your jobs
The .out file contains messages, warnings, and errors. Check it if issues arise.
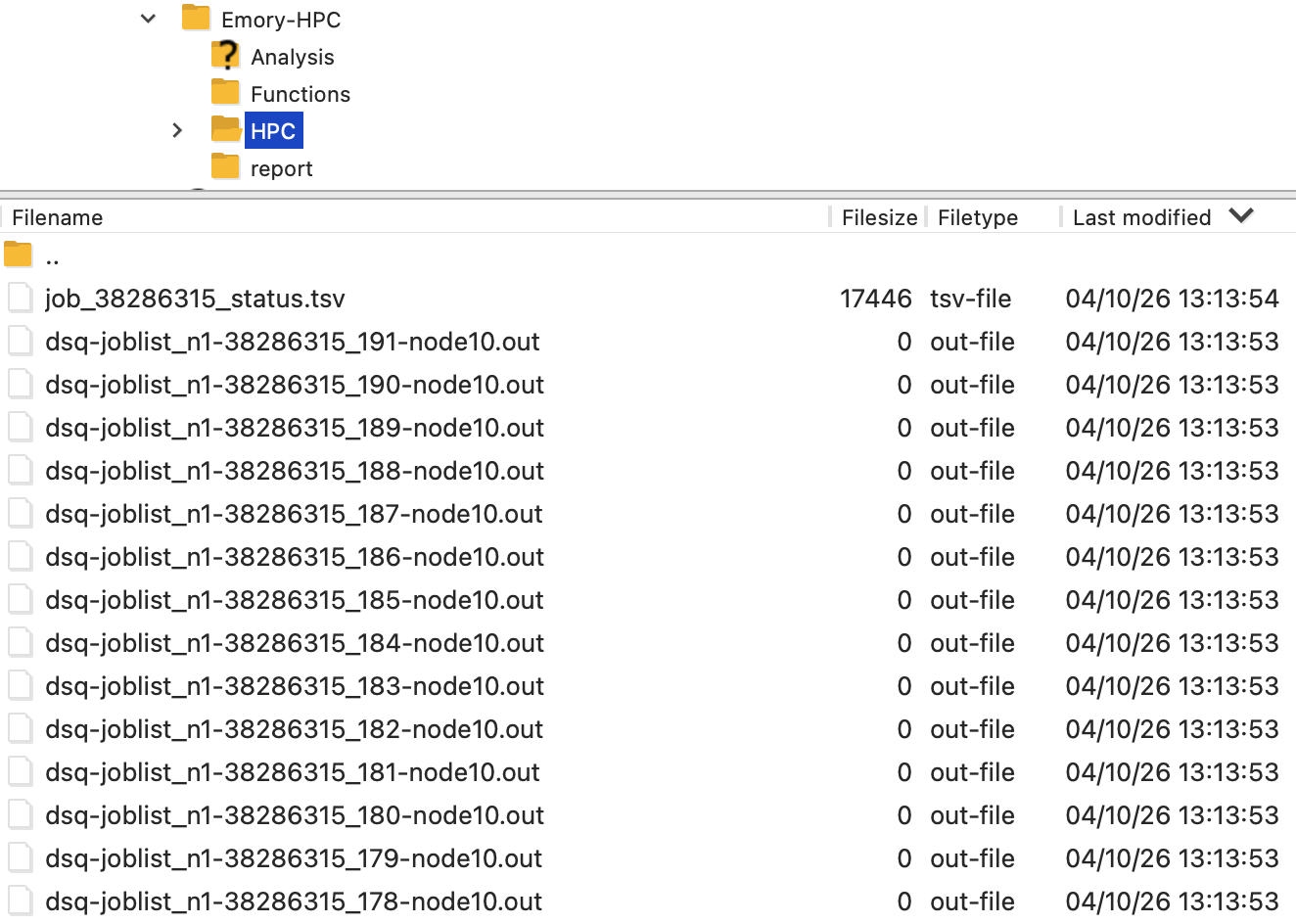
After all jobs complete successfully, you can clean up temporary files or keep them:
rm dsq* *tsv
3.4 Summarize Results
Results from each simulation are stored in HPC_Results/.
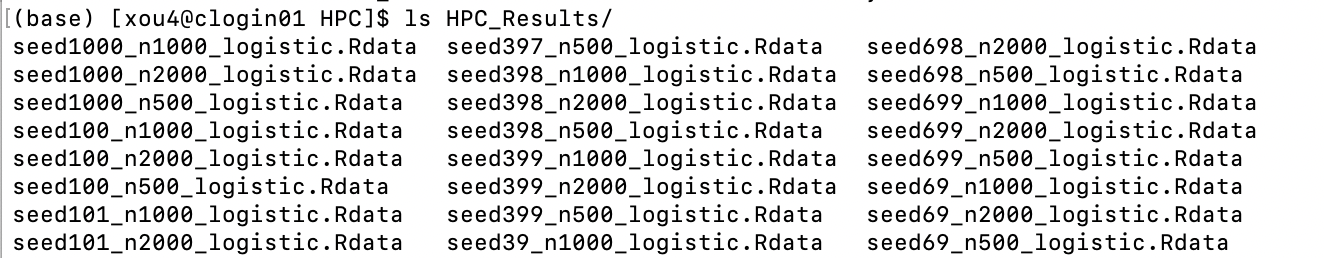
To summarize:
# make sure you are in HPC folder
Rscript 3_summarize_results.R
The file AllResults.Rdata will be saved in HPC/HPC_Results/. Transfer this file back to your local computer for further analysis.
Thanks to Anna Guo’s blog on HPC Usage for the helpful reference.